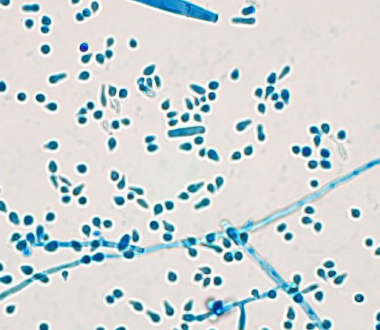
Trichophyton indotineae with macro and micro conidia
In recent years, several dermatophytes within the Trichophyton interdigitale /Trichophyton mentagrophytes Species Complex have become a significant public health concern due to increasing drug resistance and rapid transmission in human populations, leading to serious and difficult-to-treat skin infections. This complex includes 28 genotypes. Notably, T. indotineae (formerly T. mentagrophytes genotype VIII) was recently recognized as a distinct species, with resistance to the first-line drug terbinafine observed in over 50% of isolates. T. mentagrophytes genotype VII has also emerged, frequently associated with sexual transmission, and primarily reported among men who have sex with men. In addition, T. mentagrophytes genotype II has been identified in highly contagious infections affecting the scalp, hair follicles, and trunk.
Since 2022, the Wadsworth Center Mycology Laboratory has responded to the emergence of dermatophyte infections by providing identification, antifungal susceptibility testing, analysis of resistance mechanisms to the first-line drug terbinafine, and investigation of transmission patterns.
Collaborations with physicians and clinical and public health laboratorians have resulted in several publications in peer-reviewed journals
- Clinical Course, Antifungal Susceptibility, and Genomic Sequencing of Trichophyton indotineae
- Notes from the Field: Trichophyton mentagrophytes Genotype VII - New York City, April-July 2024
Notes from the Field: First Reported U.S. Cases of Tinea Caused by Trichophyton indotineae - New York City, December 2021-March 2023
Due to its specialized expertise, the Wadsworth Center Mycology Laboratory was designated by the Centers for Disease Control and Prevention as a Reference Laboratory in 2023 for the identification of emerging and drug-resistant dermatophytes through the Epidemiology and Laboratory Capacity Antibiotic Resistance Laboratory Network program for the Northeast region and the United States. As a result, the laboratory's workload has increased substantially.
To address the labor-intensive process of genotyping dermatophytes via manual alignment analysis, the laboratory developed a pipeline using a deep learning convolutional neural network model to assign Trichophyton genotypes. The team tested the model on 382 clinical isolates of Trichophyton received since May 2022 and the pipeline accurately and quickly predicted Trichophyton at both the species and genotype levels, saving time and effort. The algorithm can be implemented by any clinical, public, or reference laboratory with bioinformatics expertise. Details of the model and related findings were recently published in the April 10, 2026 edition of Journal of Clinical Microbiology https://journals.asm.org/doi/10.1128/jcm.00156-26).The work also highlights the key role of bioinformaticians at the Wadsworth Center whose skills help streamline lab workflows and deliver faster, more efficient results for patient care.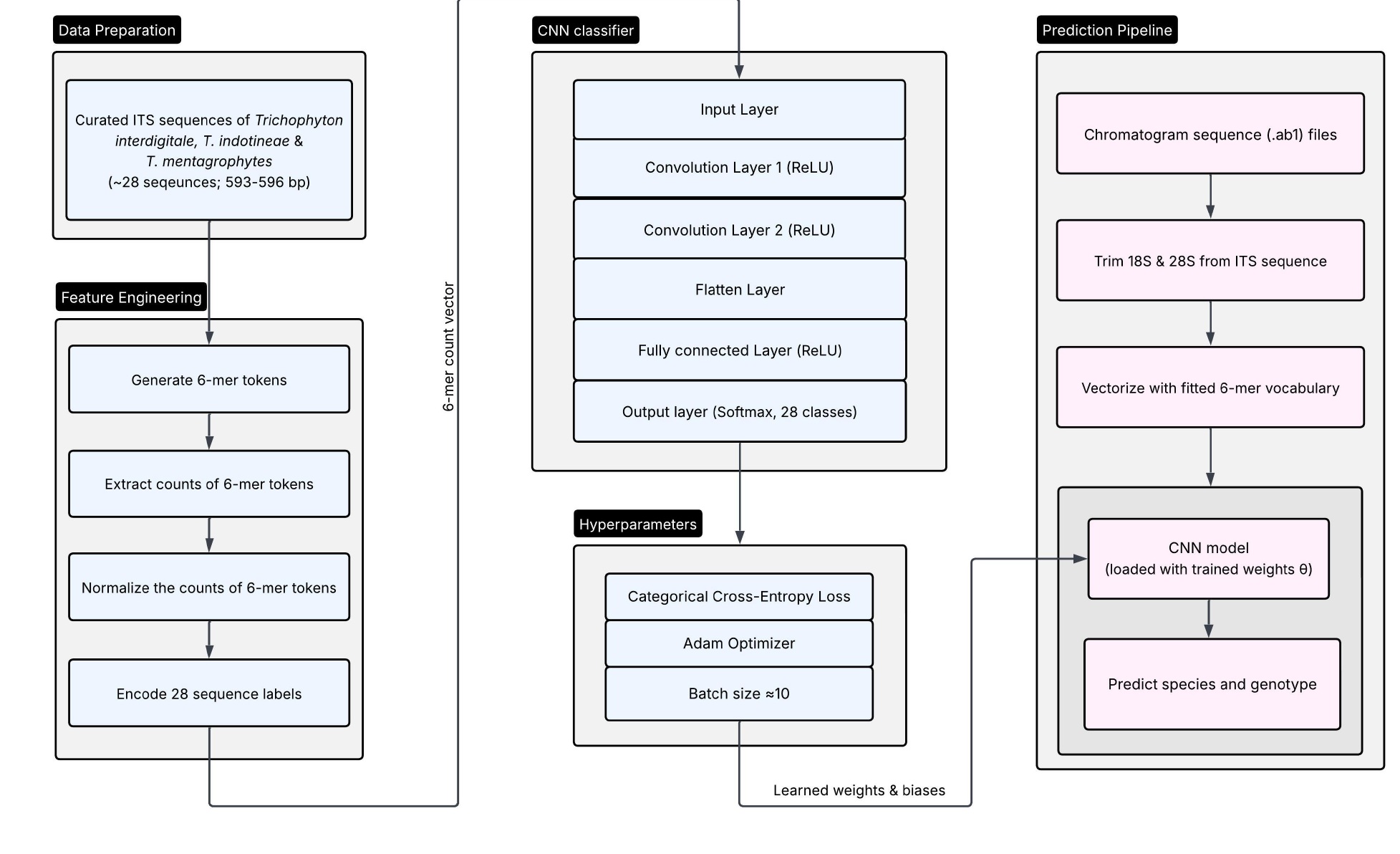
Architecture of the convolutional neural network (CNN) model based ITS sequence pipeline for the determination of genotypes within TiTmSC. The diagram illustrates the end-to-end data transformation from raw chromatogram inputs to predicted genotype.